Introduction PCR – The polymerase chain reaction (PCR) is one of the most powerful technologies in molecular biology. Using PCR, specific sequences within a DNA or cDNA template can be copied, or “amplified”, many thousand- to a million-fold using sequence-specific oligonucleotides, heat-stable DNA polymerase, and thermal cycling.
Introduction PCR
In traditional (endpoint) PCR, detection and quantification of the amplified sequence are performed at the end of the reaction after the last PCR cycle, and involve post-PCR analysis such as gel electrophoresis and image analysis.
In real-time quantitative PCR (qPCR), PCR product is measured at each cycle. By monitoring reactions during the exponential-amplification phase of the reaction, users can determine the initial quantity of the target with great precision.
PCR theoretically amplifies DNA exponentially, doubling the number of target molecules with each amplification cycle. When it was first developed, scientists reasoned that the number of cycles and the amount of PCR end-product could be used to calculate the initial quantity of genetic material by comparison with a known standard.
To address the need for robust quantification, the technique of real-time quantitative PCR was developed.
Currently, endpoint PCR is used mostly to amplify specific DNA for sequencing, cloning, and use in other molecular biology techniques.
In real-time PCR, the amount of DNA is measured after each cycle via fluorescent dyes that yield increasing fluorescent signal in direct proportion to the number of PCR product molecules (amplicons) generated.
Data collected in the exponential phase of the reaction yield quantitative information on the starting quantity of the amplification target. Fluorescent reporters used in real-time PCR include double-stranded DNA (dsDNA)–binding dyes, or dye molecules attached to PCR primers or probes that hybridize with PCR products during amplification.
The change in fluorescence over the course of the reaction is measured by an instrument that combines thermal cycling with fluorescent dye scanning capability.
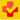
By plotting fluorescence against the cycle number, the real-time PCR instrument generates an amplification plot that represents the accumulation of product over the duration of the entire PCR reaction.
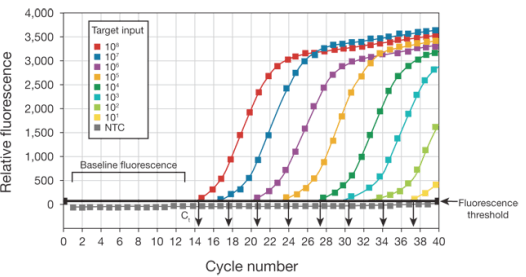
The advantages of real-time PCR include:
- Ability to monitor the progress of the PCR reaction as it occurs in real time
- Ability to precisely measure the amount of amplicon at each cycle, which allows highly accurate quantification of the amount of starting material in samples.
- An increased dynamic range of detection.
- Amplification and detection occur in a single tube, eliminating post-PCR manipulations.
Over the past several years, real-time PCR has become the leading tool for the detection and quantification of DNA or RNA.
Using these techniques, you can achieve precise detection that is accurate within a 2-fold range, with a dynamic range of input material covering 6 to 8 orders of magnitude.
Overview of real-time PCR
This section provides an overview of the steps involved in performing real-time PCR. Real-time PCR is a variation of the standard PCR technique that is commonly used to quantify DNA or RNA in a sample.
Real-time PCR steps
There are three major steps that make up each cycle in a real-time PCR reaction. Reactions are generally run for 40 cycles.
Denaturation
High-temperature incubation is used to “melt” double-stranded DNA into single strands and loosen secondary structure in single-stranded DNA.
The highest temperature that the DNA polymerase can withstand is typically used (usually 95°C). The denaturation time can be increased if template GC content is high.
Annealing
During annealing, complementary sequences have an opportunity to hybridize, so an appropriate temperature is used that is based on the calculated melting temperature (Tm) of the primers (typically 5°C below the Tm of the primer).
Extension
At 70–72°C, the activity of the DNA polymerase is optimal, and primer extension occurs at rates of up to 100 bases per second. When an amplicon in real-time PCR is small, this step is often combined with the annealing step, using 60°C as the temperature.
Two-step qRT-PCR
Two-step quantitative reverse transcriptase PCR (qRT-PCR) starts with the reverse transcription of either total RNA or poly(A) RNA into cDNA using a reverse transcriptase (RT).
This first-strand cDNA synthesis reaction can be primed using random primers, oligo(dT), or gene-specific primers (GSPs).
To give an equal representation of all targets in real-time PCR applications and to avoid the ‘3’ bias of oligo(dT) primers, many researchers use random primers or a mixture of oligo(dT) and random primers.
The temperature used for cDNA synthesis depends on the RT enzyme chosen. After reverse transcription, approximately 10% of the cDNA is transferred to a separate tube for the real-time PCR reaction.
One-step qRT-PCR
One-step qRT-PCR combines the first-strand cDNA synthesis reaction and real-time PCR reaction in the same tube, simplifying reaction setup and reducing the possibility of contamination. Gene-specific primers (GSP) are required.
READ: ELISA FAQs
This is because using oligo(dT) or random primers will generate nonspecific products in the one-step procedure and reduce the amount of product of interest.









